Processing Tab
Table of Contents
Overview
Once a data collection Run has completed with 10 or more images, the data are automatically processed using ICEflow - an SSRL automated pipeline that runs several 3rd party data processing applications that provide quick feedback on the quality of each data set.
Data processing run information and selected results are written to the Sample Database.
Once the run information appears in the database, selected information is diplayed in the Processing Tab.
If the Processing Tab is not displaying processing information:
- Make sure a spreadsheet has been assigned to the cassette in the Screening Tab
(which is currently required for writing to the Sample Database).
A default spreadsheet can be created
and assigned to the cassette.
- If you have assigned the correct spreadsheet to your cassette in the
Screening Tab and you still cannot see processing information, contact beamline support staff.
If a group does not want auto-processing results written to the Sample Database, they can opt out by checking the appropriate checkbox on the SSRL SMB Unix account request form (applies upon submittal) or by contacting their support staff. If a group opts out, no results will be recorded in our database or displayed in the Processing Tab, however auto-processing will still be carried out and the results can be found in the auto-processing directories (described below.).
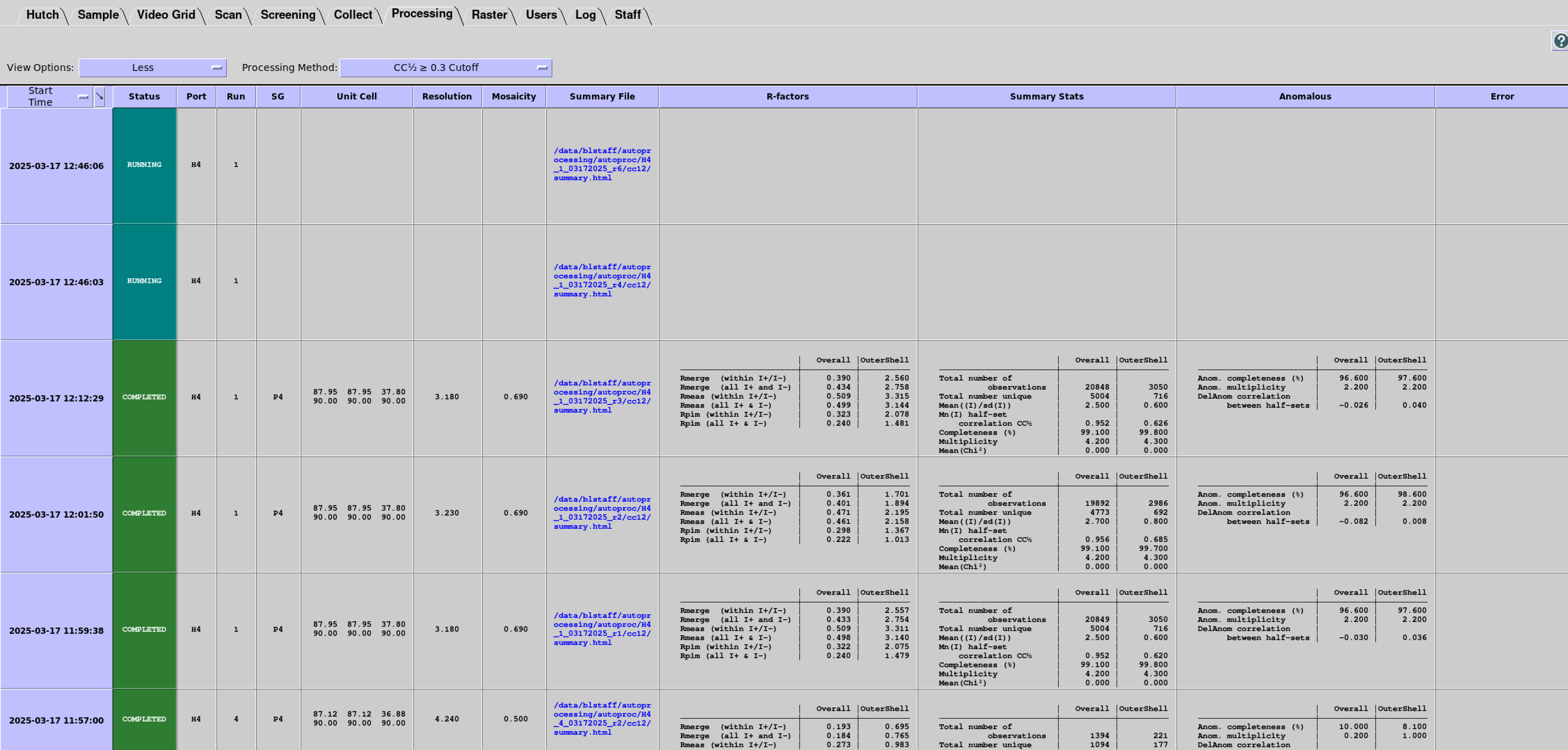
Processing Tab Layout and Navigation
Each Processing Tab row displays information and selected results for each data processing run; the table can be quickly traversed using scroll bars or arrow keys. You can use a drop-down menu to select from four Display Options: Minimum, Less, More, and All:
- Minimum - a bare-bones configuration showing essential information:
- Start Time
- Status
- Port
- Run
- SG
- Unit Cell
- Resolution
- [Resolution cutoff]
- Summary file
- Error
- Less - a configuration showing additional summary statistics:
- Mosaicity
- R-factors
- Summary Stats
- Anomalous
- More - a comprehensive list including:
- Crystal ID
- Protein
- Filename
- Image Directory
- Processing Directory
- 3rd Party Pipeline
- 3rd Party Pipeline Version
- All - adds information for staff troubleshooting:
- Sample ID
- Spreadsheet ID
- Job ID
- Hostname
The "Processing Method" drop-down menu can be used to filter the various resolution cutoff methods (i.e. CC1⁄2, I/σ(I), etc.) or to view all methods for comparison.
The first column in the table is used for sorting. The default sorting is on Start Time (when the job was submitted to the queue). Other columns can be selected for sorting by clicking on the first column header and selecting a column name in the drop down menu.
The first column can also be used to sort in the reverse direction by clicking on the arrow in the top right-hand corner of the column header.
The widths of the table columns can be changed by hovering the mouse pointer near the side of the column. When the pointer turns into a cross, click and drag the side of the column to the desired width.
The “?” button opens a webpage with documentation for the Processing Tab.
The "Status" column indicates the current status of the auto-processing job:
- Pending - the job has been added to the queue but not started yet
- Submitted - the job has been started
- Running - the job is currently in progress
- Error (highlighted in red) - the job has exited with an error. The associated error message can be found in the column labeled "Error". Processing error messages are extracted from the general log file.
- Completed - the job has finished without errors
The "Summary" column will show a path to a summary HTML file (e.g. 00_summary.html for autoPROC)
- Double-clicking on the cell with the filename will open the file as a webpage in a web browser (it may take a few seconds for the browser to open).
- Periodically refreshing the page will show the updated statistics as the processing job runs.
- The file will also display any error messages and warnings that come up.
How to Interpret Error Messages
- If an error occurs, the Status column will indicate an error and the message will be displayed in the Error column on the Processing Tab. There are two general types of errors:
- ICEflow errors (these should be labeled "ICEFLOW ERROR")
- Processing software errors (these should be labeled "AUTOPROC ERROR" for processing with autoPROC)
- If you see ICEflow errors, please contact your beamline staff.
- autoPROC errors most often reflect issues with processing the data; some of the most common types are:
- indexing errors by XDS
- integration errors by XDS
- scaling errors in apScale or XSCALE
ICEflow extracts autoPROC error messages from the "top" log file (typically named out-{cutoff}.log);
these errors most often point to the log files for specific processes (e.g. indexing) and provide relative paths to them.
Inspect these logs if you need more detailed information about what went wrong.
The autoPROC
manual lists a few common errors that can be encountered when running autoPROC
as well as a few general suggestions for how to handle them. ICEflow is designed to avoid the more basic errors (for example,
all SSRL beamline-specific settings have been implemented already), but if any of these errors crop up, please let the beamline staff know.
Viewing Processing Results from Previous Spreadsheets
A standalone Processing Tab can be opened to view processing results from other spreadsheets by clicking on "Previous Spreadsheets". When the tab first opens, a list of all spreadsheets associated with the account will be displayed in the Spreadsheet View.

The Spreadsheet View currently displays the Upload Time, Spreadsheet Name, Spreadsheet ID and the Number of Datasets associated with a particular spreadsheet. If the selected spreadsheet is currently assigned to a beamline, the last two colums, Assigned Beamline and Assigned Position, will indicate the assigned beamline and the position of the cassette in the robot dewer. To view the processing runs for a particular spreadsheet, highlight a row and click the "View Processing Results" button or simply double click on the row. This will open the associated Results View in the standalone Processing Tab. The Results and Spreadsheet Views can be toggled using the same button.
The standalone Processing Tab can also be accessed for viewing past results when the user is offline. Simply click on the Blu-Ice starter icon in the linux desktop and click on "Processing Tab".
Where Are My Data Located?
The data processing directories can be quickly accessed by clicking on the link listed in the Processing Directory column using the View Options "More" or "All".
For those that have opted out, a symbolic link to the processing directories can be found in the image directory, for example:
/data/{username}/mb_test/A5/autoprocessing_autoproc_8c562b
How to Modify the Processing Script and Reprocess Datasets Manually
- Click on the Processing Directory link.
- Make a copy of the processing script:
> cp run-{cutoff}.sh my_new_run.sh
- Open the new script in the geany or gvim text editor and modify the autoPROC launch string as needed:
> geany run-{cutoff}.sh
CRITICAL - make sure to change the folder name after the -d argument or the script will not run! e.g.:
process [pre-existing arguments] -d new_processing_folder [rest of arguments]
- Save the new version and run my_new_run.sh:
> ./my_new_run.sh
ICEflow Pipeline (autoPROC)
The intial version of ICEflow, deployed autoPROC v1.0.5 in a default configuration.
Changes made to the pipeline, 3rd party software, configurations, input parameters, etc.
are listed and documented in the next section.
Resolution Cutoffs
- cc12 – Data are processed using a resolution cutoff corresponding to a value of CC1⁄2 that is dynamically calculated by autoPROC.
- isigi - Data are processed using a resolution cutoff corresponding to a value of I/σ(I) that is calculated dynamically by autoPROC.
- nocutoff - Data are processed without using a resolution cutoff, i.e. to the corners of the detector.
Programs and Output
- autoPROC - the data processing pipeline. The general log files are written into the top directory:
XDS – performs indexing, refinement, and integration.
The input file XDS.INP - generated automatically by autoPROC - supplies the default parameters to the program
and is based upon information stored in the header of the diffraction images (detector type and distance, oscillation start and range, number of images in the date set, etc.).
The important output files from XDS can be found in the cutoff subfolders, which contain:
IDXREF.LP - the results of the automated indexing to find the unit cell parameters and an idea of what the crystal symmetry is.INTEGRATE.LP - the full log of the processing.CORRECT.LP - gives an indication of the data quality and resolution.XDS_ASCII.HKL - contains integrated intensities.
POINTLESS – is run often during the workflow to analyzes the data for twinning, symmetry, and will identify the correct space group.
AIMLESS - takes the output from POINTLESS, calculates scale factors between all the images in the data set, applies the scales, and merges all the reflection data together to give an output file containing one copy of each reflection (the unique data set). While the key output from AIMLESS is included in the general autoPROC output file; the full log can be found in the {cutoff} directory.
CTruncate - reads the output from AIMLESS and attempts to put the data onto an absolute scale and generates structure factor amplitudes (F) from the reflection intensities (I). Its output can be found in the {cutoff} directory
References
- AUTOPROC - Vonrhein, C., Flensburg, C., Keller, P., Sharff, A., Smart, O., Paciorek, W., Womack, T. & Bricogne, G. Data processing and analysis with the autoPROC toolbox. Acta Crystallographica D67, 293-302 (2011).
- AUTOPROC - Vonrhein, C., Flensburg, C., Keller, P., Fogh, R., Sharff, A., Tickle, I.J. and Bricogne, G., Advanced exploitation of unmerged reflection data during processing and refinement with autoPROC and BUSTER. Acta Crystallographica D80(3) (2024).
- XDS - Kabsch, W. XDS. Acta Crystallographica D66, 125-132 (2010).
- POINTLESS - Evans, P.R. Scaling and assessment of data quality, Acta Crystallographica D62, 72-82 (2006).
- AIMLESS - Evans, P.R. and Murshudov, G.N. How good are my data and what is the resolution? Acta Crystallographica D69, 1204–1214 (2013).
ICEflow Versions and Release Notes
- ICEflow-1.5.0 - released on 03/19/2025
- ICEflow jobs are now submitted to a queue using the Slurm workload manager.
- Processing job submission parameters tweaked for optimal resource usage.
- Major changes to the GUI:
- A new status - "Pending" - appears for jobs waiting to run.
- New columns added to the Layout, with two settings available to users.
- A clickable link to a summary webpage file added.
- autoPROC log parsing takes into account new formatting.
- ICEflow-1.4.0 - released on 01/14/2025
- Added data collection strategy calculation (iMosflm) to ICEflow packages
- Strategy can be found on the Collect Tab
- ICEflow-1.3.0 (autopPROC) - released on 09/12/2024
- Added error extraction from logs and reporting in the Processing Tab.
- Reorganized result reporting to database and UI.
- ICEflow-1.2.1 (patch) (autopPROC) - released on 06/21/2024
- Fixed issue where autoPROC could not index images collected with a vertical offset on a Pilatus detector.
- Changed symlinks in data folder to point to the top processing folder.
- ICEflow-1.2.0 (autoPROC) - released on 06/18/2024
- Changed to a more descriptive convention for output folder:
/data/{username}/{3rd_party_software}/{filename_prefix}_{run_number}_{date}_{time}/{cutoff}
- The
README file now contains explicit path to source data and image file template, for easier reference.
- Documentation below edited to reflect these changes.
- ICEflow-1.1.0 (autoPROC) - released on 06/12/2024
- Fixed an issue where both cutoff versions output data with a CC1/2-based resolution cutoff; the I/σ(I)-based cutoff is now enabled.
- Added a no-cutoff option as documented below.
- The summary.html files are now copied to the top folder for easier access.
- ICEflow-1.0.0 (autoPROC) - released on 06/05/2024
- Initial release, with only autoPROC pipeline enabled.
Pipelines Running before the Inception of ICEflow (before 6/5/2024)
For the SSRL pipeline (and the xia2 test pipeline),
look in the image file directory for the symbolic link to the directory with the processed data (these links will all begin with 'autoprocessing').
The processing directory contains the README file that describes the pipeline used for data processing.
If you can't find what you're looking for, contact your user support staff member.
More information on the software supported by the SSRL-SMB Macromolecular Crystallography division is available on our software webpage.
|



